This protocol helps to set up a Wound Healing Assay using HoloMonitor® M4 and the HoloMonitor® App Suite software. The HoloMonitor® Wound Healing Assay provides an automated label-free wound healing assay, measuring gap closure. Additionally, selected cells can be individual cell tracked for detailed individual cell movement and morphology analysis.
Requirements:
- HoloMonitor® M4, placed in incubator
- HoloMonitor® M4 App Suite
- Culture vessel of choice with cells
- HoloLid™ for selected vessel
- Vessel holder for selected vessel
Output:
- Gap width (μm)
- Cell covered area (μm2 and %)
- Cell free area (μm2 and %)
Reanalysis:
- Guided assays
- In-depth analysis
PREPARATIONS
Materials:
- HoloMonitor® M4, placed inside the incubator
- HoloMonitor® App Suite software
- Cell culture vessel. Please check our list of recommended vessels.
- HoloLid™ for the selected vessel
- Vessel holder for selected vessel
- Setup and Operation Manual for HoloMonitor® M4
► We recommend using ibidi® µ-Dish 35 mm with Culture-Insert 2 Well high (cat. # 81176) or ibidi® µ-Plate 24-well with Culture-Insert 2 Well (cat. # 80241)
Steps
Seed the cells to at least 90 % confluence in the inserts according to the ibidi protocol.
Place the vessel in the incubator and let cells attach for 2-24 hours.
Start the software and wait for complete instrument initialization.
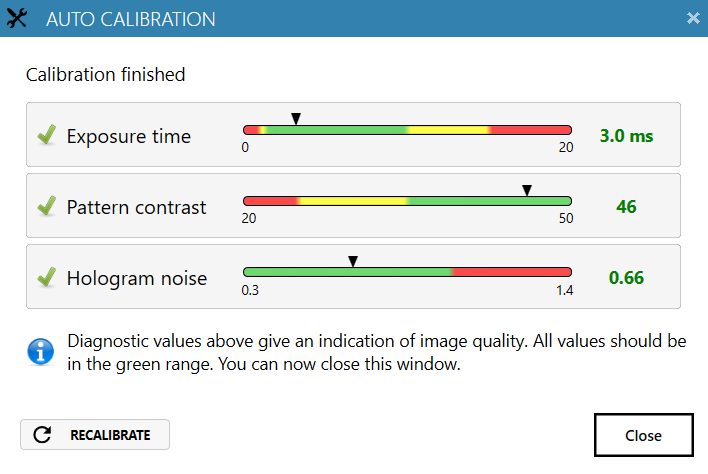
Run an auto-calibration. With successful calibration (as shown in the picture), the instrument is ready to use.
Sterilize the HoloLids™ according to the HoloLid™ sterilization and use protocol.
Add the treatment to your cells. The final working volumes per well, essential for using HoloLids™, are shown in the table above.
Slide the cell culture vessel onto the Vessel holder, its grips facing towards you. Ensure that the vessel is parallel to the holder. There is a spring that holds the vessel in place.
►When using multi-well plates, place them with the cut-off corner to the left.
Replace the standard lids with the HoloLid™.
Put the vessel holder with the sample on the HoloMonitor® M4 stage.
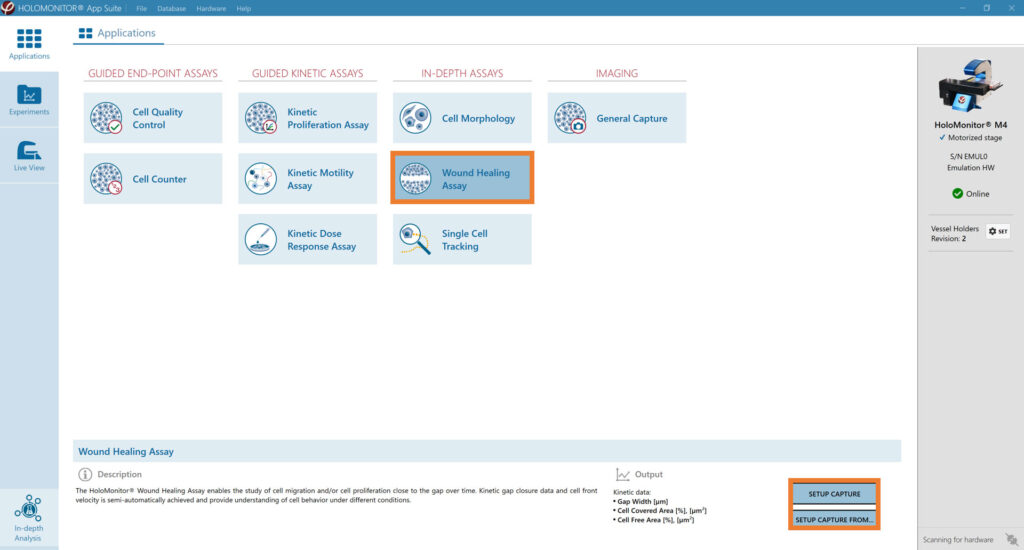
Select the Wound Healing Assay and proceed by clicking the Setup Capture button.
EXPERIMENT SETUP
Basic setup: Describe the experiment and assign treatments to the wells
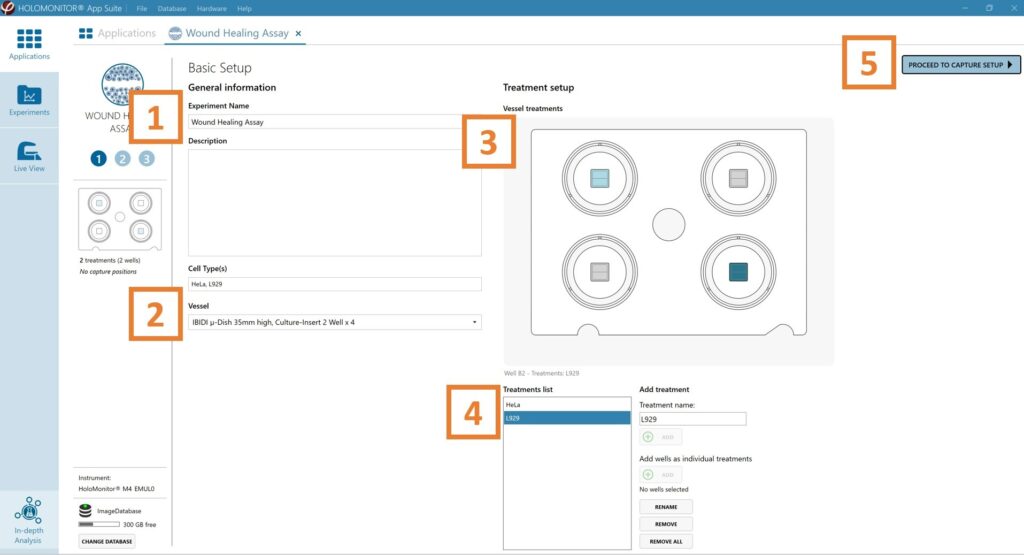
Enter the experiment name, optional experiment
description and cell types.
Select the correct vessel map from the drop-down list.
Map treatments and conditions on the vessel map. Select wells by marking them with the left mouse button while moving the cursor over the well/s.
Add the treatment name/s in the text box below the vessel map and click Add/ press Enter. Marked well/s are light blue, selected wells will appear dark blue.
Proceed to Capture setup.
Capture setup: Select the experiment time settings and choose capture positions
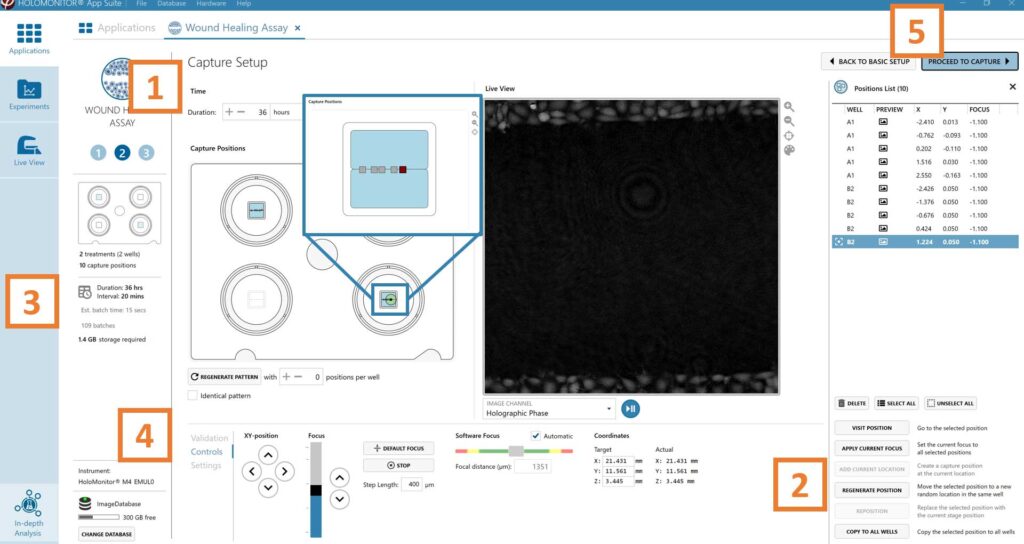
Adjust the default settings for duration and interval.
Add capture positions: The position list is open by default. Click positions on the vessel map and add them to the position list with the Add current location button. In case the image quality is poor, a warning sign appears. Adjust focus or position location if necessary.
►Note that the gap might not be exactly where the vessel map indicates.
Ensure that the storage requirement for the experiment does not exceed the computer capacity.
► When running an experiment, data needs to be stored on the computer connected to the instrument. Storing data on an external drive (e.g. connected via USB or internet server) may cause data loss due to erratic USB connections or poor internet connection.
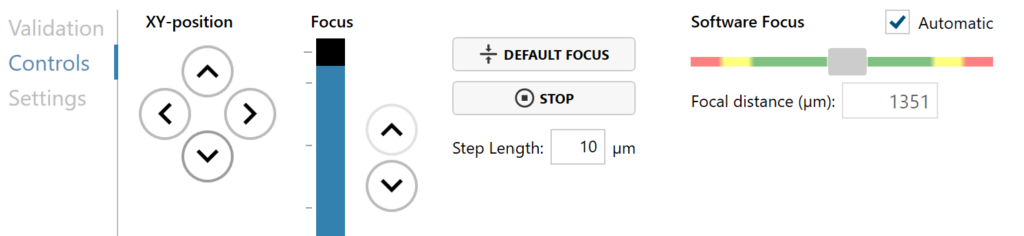
Run a full or quick validation of the selected positions to ensure good image quality.
When satisfied with the experiment setup, click Proceed to Capture.
►If there are no positions with both wound edges visible at the same time, add two positions that are parallel to each other and have one edge with cells. Use XY position Arrow buttons to move the stage in steps and when satisfied, press Add Current Location button.
Capture: Review the experiment in real-time during the time-lapse
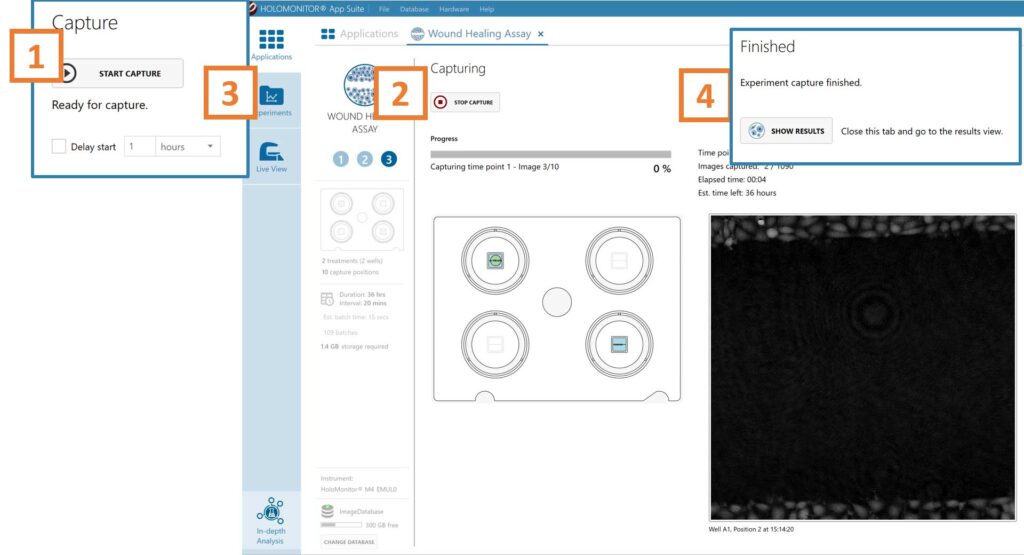
Click Start Capture.
To stop the experiment ahead of time, click the stop button.
► Note that it is NOT possible to restart the experiment once it has been stopped.
Go to the Experiments tab and open your ongoing experiment to preview the results during the run.
► Wait for the experiment to finish before starting In-depth Analysis.
When the Experiment capture finishes, click the Show Result button to get directly to the Results page.
RESULTS & ANALYSIS
Experiments tab
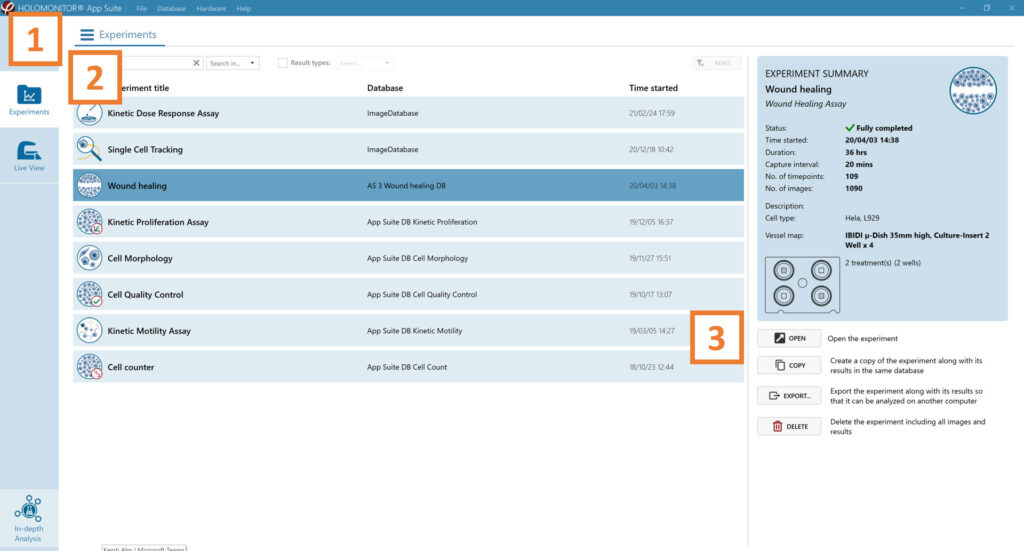
Click Experiments to see a list of the experiments.
Click on the experiment title to open an experiment summary.
Click Open to open the results page.
Experiment overview tab
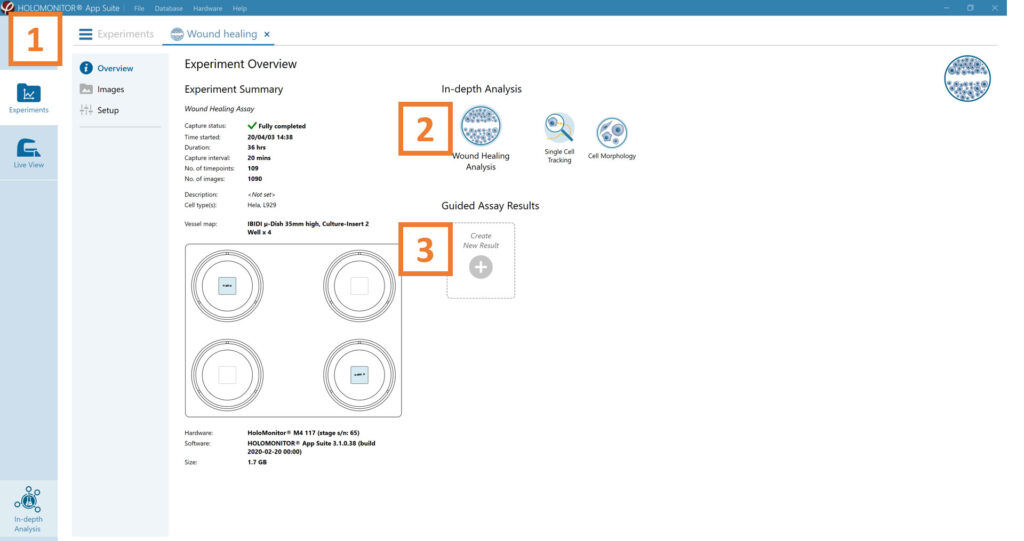
See the experiment summary, view all images, and go to the experiment setup by choosing the respective tab.
Generate in-depth analysis data from the captured images by clicking on the Wound Healing icon. A new window for the in-depth analysis will open.
Create New Guided Assay Results from this experiment by clicking the respective button.
In-depth analysis —Wound healing tab
Begin by adding frames to the analysis either by drag and drop or using the Add selected or Add all button.
►Check the image quality before using it for in-depth analysis. See the Image quality guide for more information.
Adjust the image threshold and object size and click Apply to all frames. You can alter the viewing options.
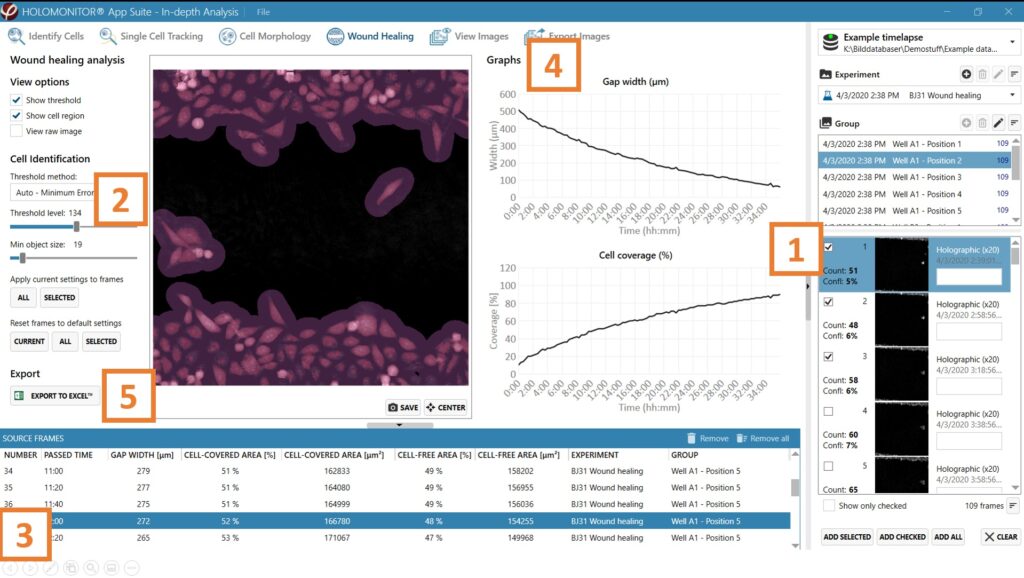
Based on the identification in the previous step, result values are generated and displayed in the table: gap width (mm), cell-covered area (% and μm2), and cell-free area (% and μm2).
►In case of outliers, select the outlier frame in the list and adjust the threshold for that specific frame.
Check the gap width and cell coverage graphs to ensure that the values decrease evenly with time.
Export to Excel for further analysis. The exported data include gap width (μm), cell-covered area (% and μm2), and cell-free area (% and μm2), graphs for gap width, cell coverage, and the settings overview.
►From the results, it is easy to get the speed of the cell front (cell front velocity).
►If there was only one edge with cells visible in a field of view. To evaluate gap closure kinetics, calculate cell front velocity for each edge with cells and assess cell covered area (%) change in the field of view.
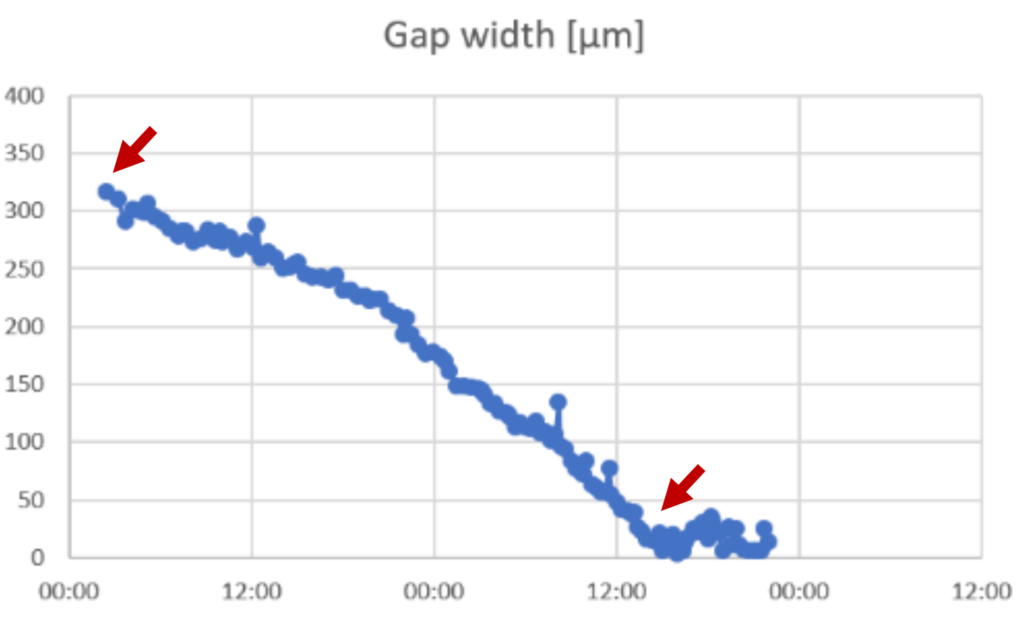
►Calculate the speed of the cell front by subtracting the last value for gap width in the linear phase from the first value for gap width within the linear phase and divide with the number of hours between the values.
►In this example: (321-25) μm/36 h = 8 μm/h
This can also be performed in the Excel datasheet.
►Note that it is important to use only data from the linear phase of the slope.
One experiment — multiple results
- This section helps to reanalyze data between different assays using HoloMonitor® App Suite software.
Generating In-depth Assay results
Generating Guided Assay results
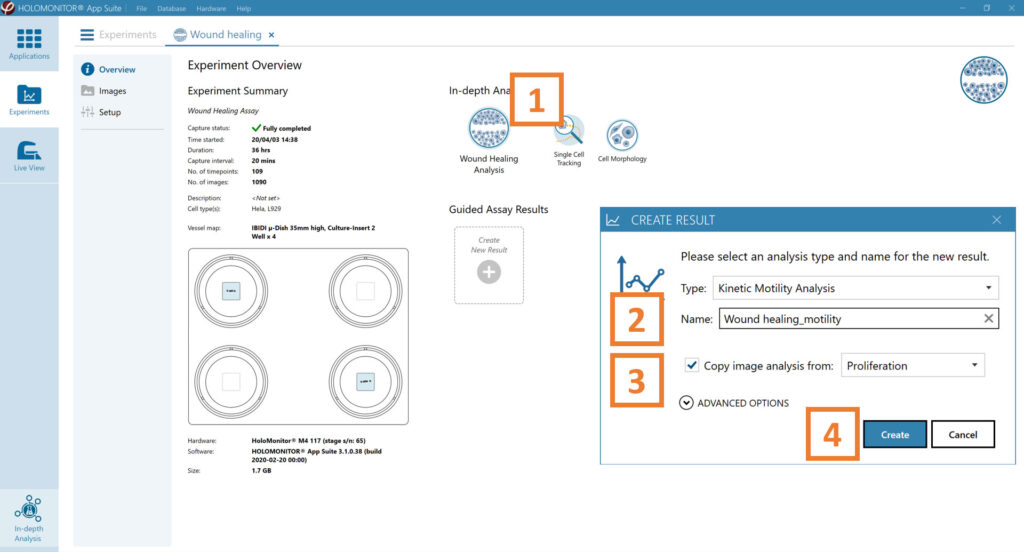
In the Experiment overview page under Guided Assay Results select Create New Result.
Choose the type of analysis in the pop-up window and name the new result.
Tick copy image analysis from and select the experiment to copy from. This will copy the image analysis settings from the selected result including all changes.
► For further data analysis steps, please see the respective assay protocol.
Press create.