created using HoloMonitor®
Time-lapse Imaging Cytometry
Time-lapse imaging cytometry by holographic microscopy allows non-invasive visualization and analysis of live cell populations by tracking and quantifying individual cells.
Understanding Cellular Dynamics
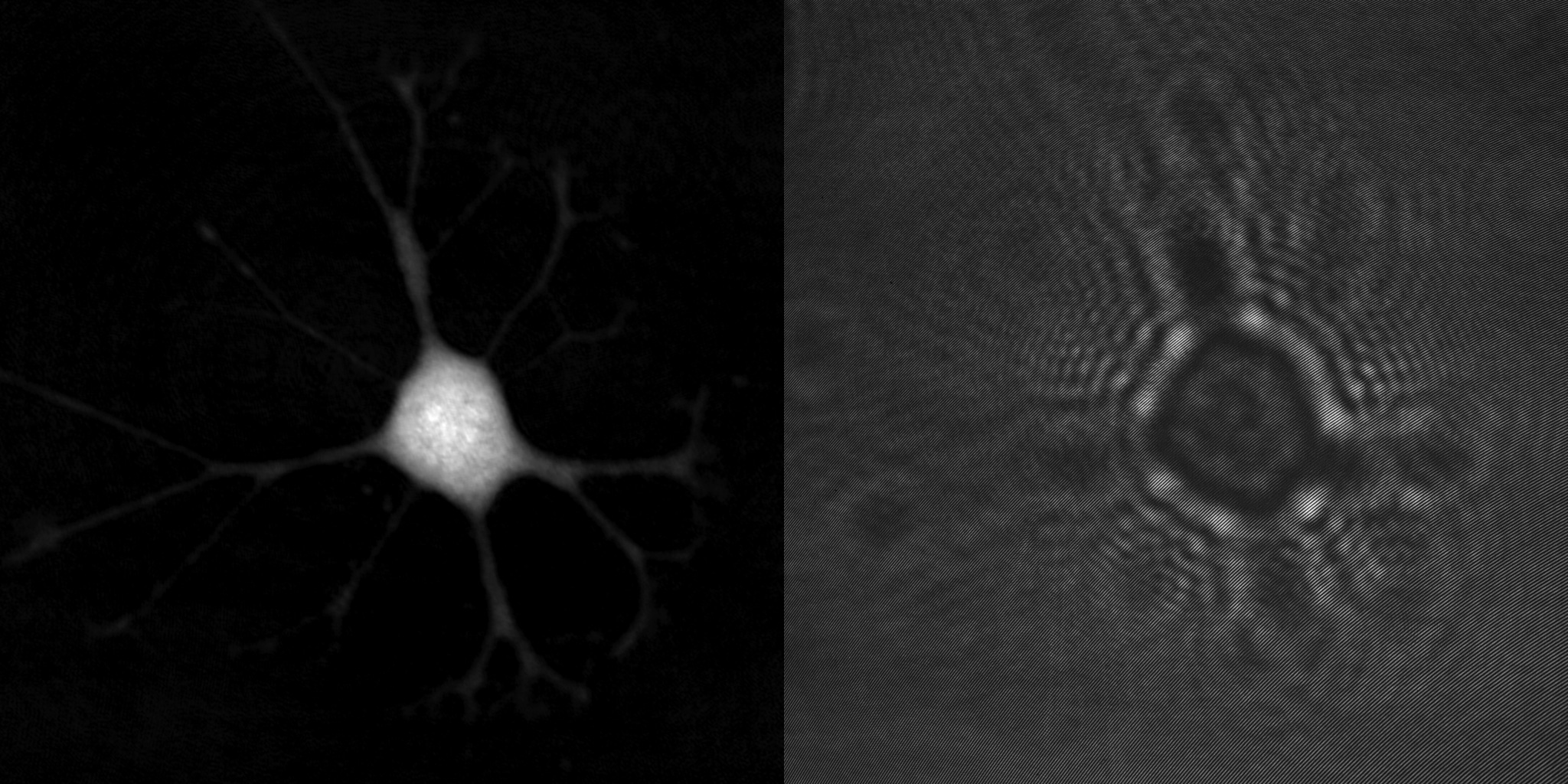
An advantage of holographic microscopy is that the created quantitative phase images are focused when viewed, not when recorded. This makes the HoloMonitor time-lapse cytometer ideal for long-term imaging and analysis of living cells by means of time-lapse microscopy, which acquires a series of cell images at regular time intervals to analyze the dynamics of various cellular events. Unfocused images, caused by focus drift, are simply refocused by letting the computer software recreate the phase image from the recorded hologram.
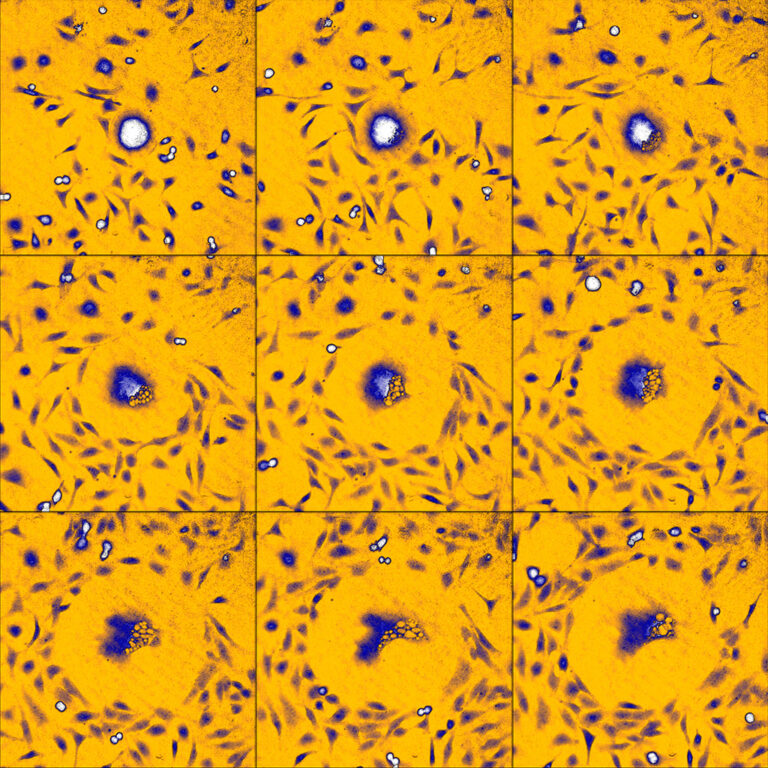
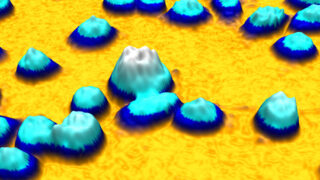
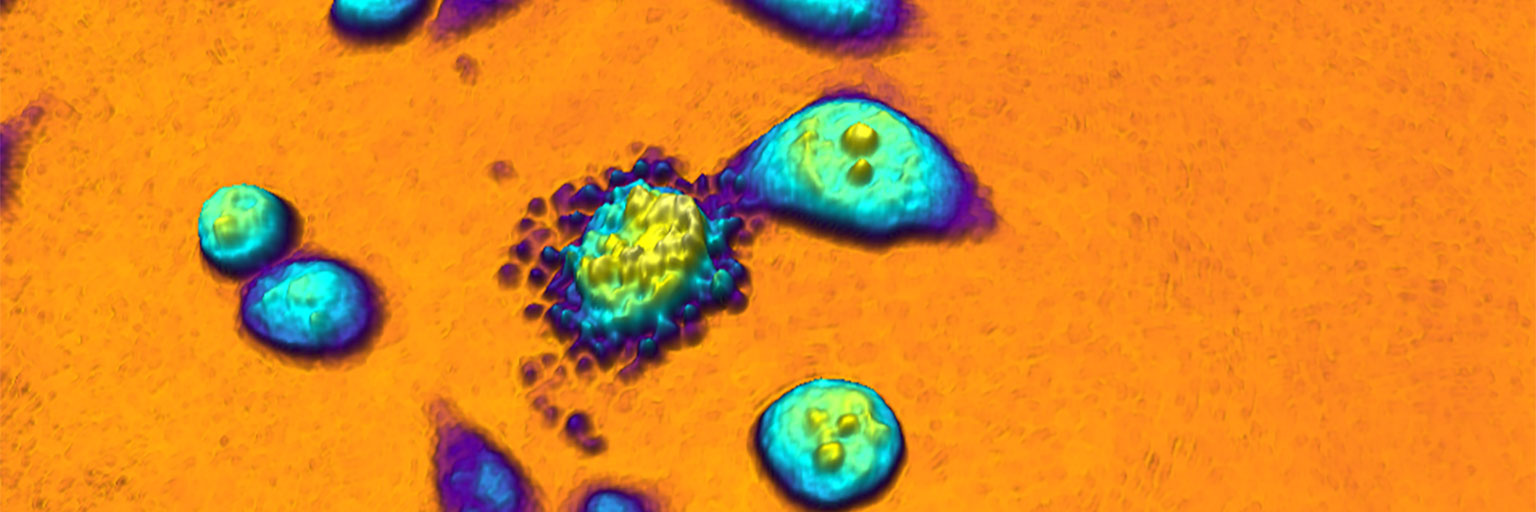
An example time-lapse image sequence of a monster HeLa cell, imaged by HoloMonitor. HeLa cells are cancer cells named after Henrietta Lacks, who in the early 1950s donated the first cells that was successfully kept alive and cultured in a laboratory environment. HeLa cells and other immortal cells are today routinely cultured by scientists to study the complex behavior of cells and their response to drug treatments.
In addition to identifying each individual cell, HoloMonitor provides data for analysis of more than 30 morphological parameters. However, the true power of time-lapse cytometry first emerges when the same cells are monitored over time. The HoloMonitor design utilizes recent technological advances to allow time-lapse image sequences of cultured cells to be effortlessly recorded over long time periods.
With HoloMonitor installed in a cell incubator, the cells are kept in a cell friendly environment during the entire experiment. Long-term live cell kinetic data can easily be obtained using time-lapse microscopy. Images are recorded at selected intervals, down to 1 image/sec. Depending on the application, cell images are played back as a video recording to aid analysis of dynamic cell behavior.
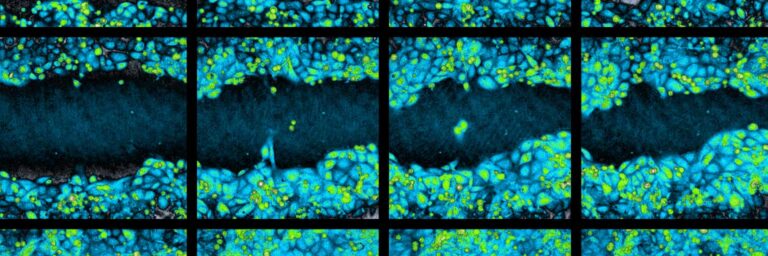
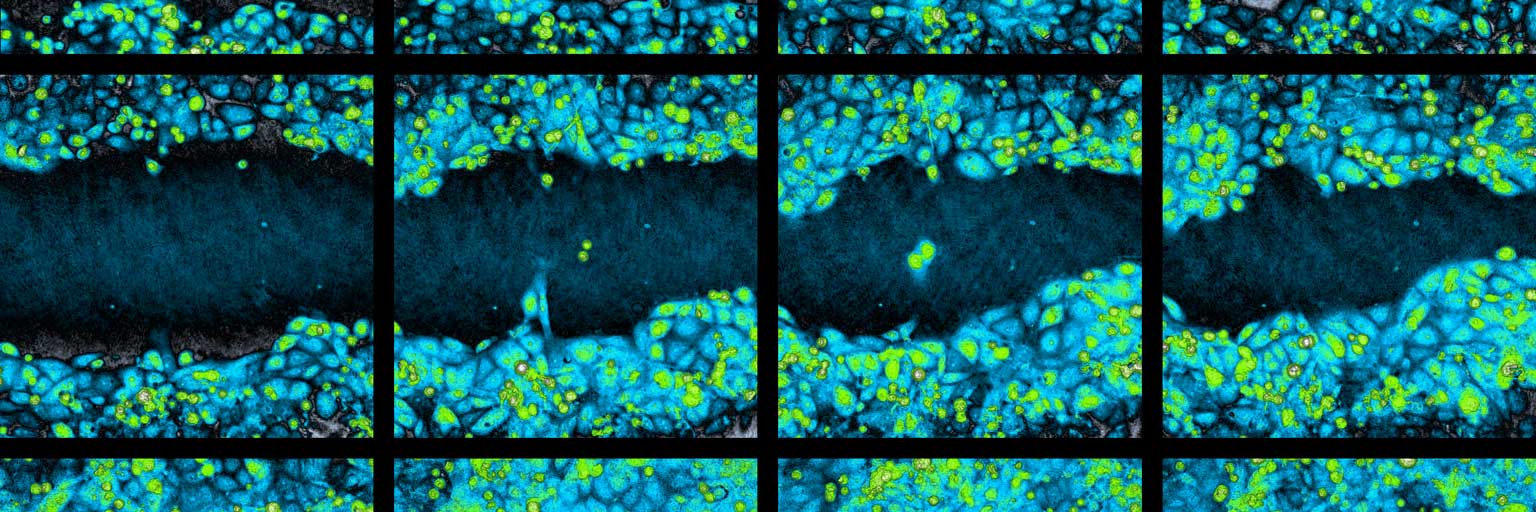
Time-lapse image sequence of a wound healing assay experiment.
Cytometric Imaging Software
From recorded time-lapse image sequences, the HoloMonitor App Suite software helps the user to automatically extract and kinetically analyze live cell population data based on individual cell data. When preferred, individual cell data — such as cell count, cell morphology, cell velocity, and cell division rate — can all be measured from the same experimental time-lapse data, without requiring additional experiments.
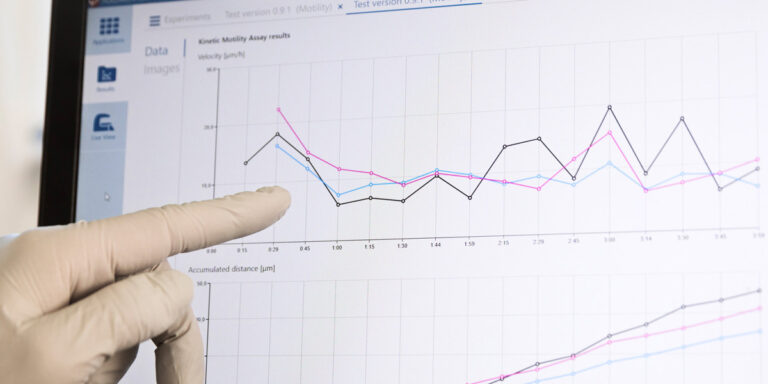
HoloMonitor App Suite
Time-lapse Cytometry References

Moving into a New Dimension: Tracking Migrating Cells with Digital Holographic Cytometry in 3D
Authors: Anette Görloff Wingren
Journal: Cytometry Part A (2018)
Research Areas: Cancer research
Keywords: HoloMonitor M4, cell motility, cell migration, cell morphology, cell tracking, cytometry, digital holography, migration, motility, quantitative phase imaging, scratch assay

HoloMonitor M4: holographic imaging cytometer for real-time kinetic label-free live-cell analysis of adherent cells
Authors: Mikael Sebesta et al.
Journal: Proceedings, Quantitative Phase Imaging II (2016)
Research Areas: Method development
Keywords: imaging cytometer, label-free, time-lapse, holomonitor m4, cytometric analysis software, incubator environment, live-cell analysis, digital holographic technology